Published
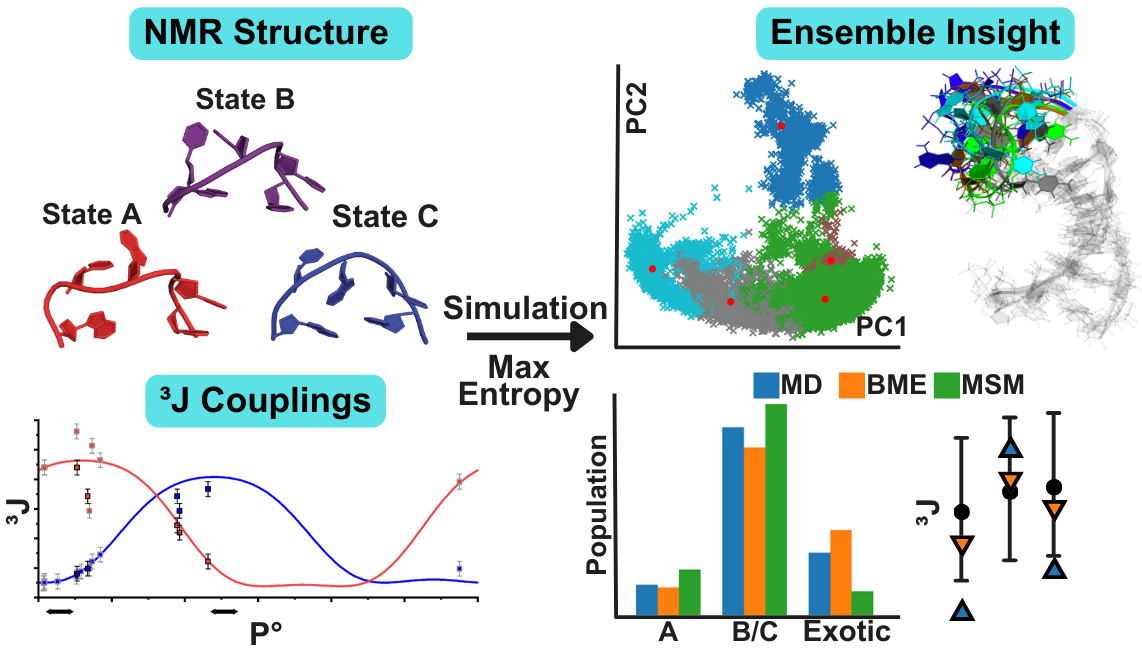
Integrated NMR and MD structure and dynamics of the stem loop II motif (s2m) from the Omicron variant of SARS-CoV-2
Tobias Matzel#, Joseph A. Makowski#, Adam H. Kensinger, Andreas Oxenfarth, Maria A. Wirtz Martin, Jeffrey D. Evanseck, Harald Schwalbe
Abstract
The stem-loop-II motif (s2m) is a conserved viral RNA element located in the 3′UTR of different viruses including SARS-CoV-2. High resolution 3D structural data for s2m are only available for the fundamentally different SCoV-1 version and difficult to access for SARS-CoV-2 due to the highly dynamic nature of the s2m RNA element. With the Omicron variant, a large deletion occurred for s2m resulting in a relatively short hairpin with an apical pentaloop. We determined the NMR solution structure of s2mOmicron using a variety of torsion-angle sensitive NMR parameters in addition to NOE distance restraints. Surprisingly, relatively high {1H},13C heteronuclear NOE values, averaged ribose 3JHH-coupling constants (H1′H2′; H3′H4′) and dipole(H1′-C1′), dipole(H6/8-C6/8)-CCRs hinted towards significant dynamics for the small pentaloop making structure calculations solely relying on NMR data insufficient. To address this problem, we performed ten 1 microsecond MD simulations from the NMR structure bundle as a starting point and applied Bayesian Maximum Entropy reweighting to refine the ensemble with the 3J-coupling constant data. Our results from the combined methodology provide a detailed view of the conformational dynamics of the Omicron variant of s2m characterized by different stacking patterns, ribose repuckering and overall heterogeneity of the torsion angles for the loop nucleotides.
DOI and authorship note
https://doi.org/10.1261/rna.080576.125
# Shared first authorship.
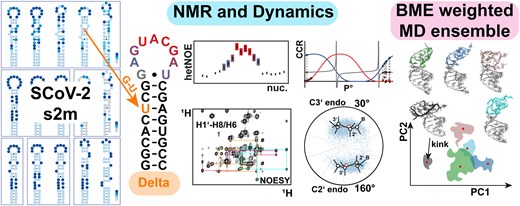
Structural heterogeneity and dynamics in the apical stem loop of s2m from SARS-CoV-2 Delta by an integrative NMR spectroscopy and MD simulation approach
Maria A. Wirtz Martin#, Joseph A. Makowski#, Tobias Matzel, Adam H. Kensinger, Alexander Herr, Christian Richter, Hendrik R. A. Jonker, Anna Wacker, Jeffrey D. Evanseck, Harald Schwalbe
Abstract
In structured RNAs, helical elements are often capped by apical loops that are integral structural elements, ranging from 3 to >20 nts of size on average, and display a highly heterogeneous energy landscape profile, rendering structural characterization particularly challenging. We here provide a characterization of the SARS-CoV-2 Delta s2m element containing a highly dynamic nonaloop using an integrative approach of nuclear magnetic resonance spectroscopy, small angle X-ray scattering, and molecular dynamics simulations. We further explored the conformational space in the s2m nonaloop and its transient closing 5′-G-U-3′ base pair by MD simulations weighted by experimental NMR observables, leading to a comprehensive representation of the s2m nonaloop motif. Our deconvolution of the ensemble into conformations and dynamics provides a basis for future ensemble-functional characterization of RNA structures featuring dynamic motifs.
DOI and authorship note
https://doi.org/10.1093/nar/gkaf552
# Shared first authorship.
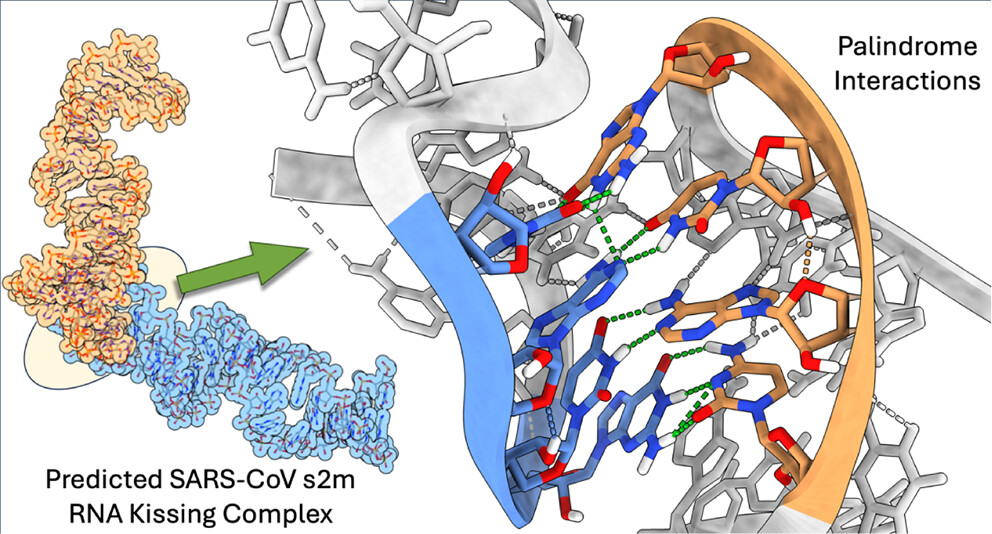
Structural Prediction of Coronavirus s2m Kissing Complexes and Extended Duplexes
Adam H. Kensinger, Joseph A. Makowski, Mihaela Rita Mihailescu, Jeffrey D. Evanseck
Abstract
The three-dimensional atomistic-resolution structure and dynamics of RNA kissing complexes and extended duplexes, homodimers formed through palindromic base pairing, are crucial for understanding viral replication and structure-informed therapeutic design. In this work, we evaluate the VFold3D/LA-IsRNA pipeline for resolving 3D structures of s2m kissing complexes and extended duplexes by validating its accuracy with blind and referenced predictions against experimental HIV-1 DIS structures. Following MD refinement, the SARS-CoV s2m kissing complex adopts stacking palindromic basepair triplets, whereas SARS-CoV-2 and Delta s2m only form canonical palindrome basepairs, explaining their relative dimer instability suggested by PAGE band intensity. Overall, the predicted structures provide an atomistic foundation for data-driven design of antiviral therapies to disrupt viral lifecycle or immune-response-relevant interactions.
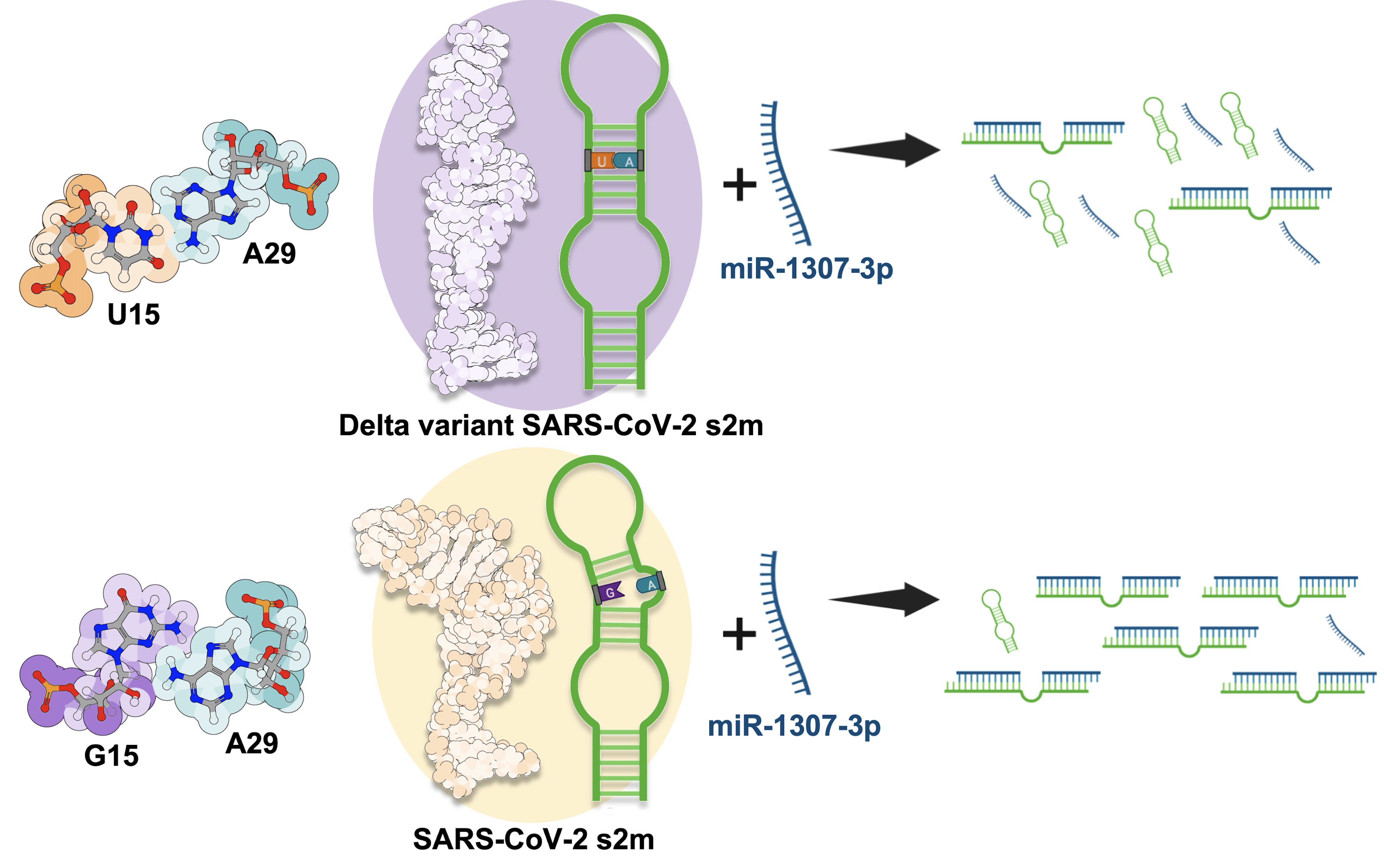
Effect of the SARS-CoV-2 Delta-associated G15U mutation on the s2m element dimerization and its interactions with miR-1307-3p
Caylee L. Cunningham, Caleb J. Frye, Joseph A. Makowski, Adam H. Kensinger, Morgan Shine, Ella J. Milback, Patrick E. Lackey, Jeffrey D. Evanseck, Mihaela-Rita Mihailescu
Abstract
The stem loop 2 motif is a highly conserved 41-nucleotide hairpin structure in the SARS-CoV-2 genome and a potential therapeutic target. This work identifies the changes between Delta and earlier SARS-CoV-2 s2m elements and characterizes how those changes affect dimerization and interactions with host microRNA miR-1307-3p. Based on NMR spectroscopy assignments, we show that the U15-A29 base pair closes, stabilizing the upper stem without overall secondary-structure deviation. Increased stability of the upper stem did not affect conversion of kissing dimers into a stable duplex conformation, but the G15U mutation drastically impacted host microRNA binding. These findings demonstrate that the observed mutation alters s2m secondary structure with downstream consequences for viral interactions with host regulatory RNA.
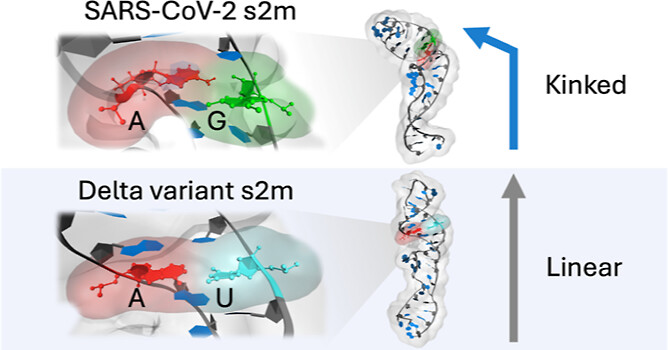
Delta SARS-CoV-2 s2m Structure, Dynamics, and Entropy: Consequences of the G15U Mutation
Joseph A. Makowski#, Adam H. Kensinger#, Caylee L. Cunningham, Caleb J. Frye, Morgan Shine, Patrick E. Lackey, Mihaela Rita Mihailescu, Jeffrey D. Evanseck*
Abstract
Bioinformatic analysis of the Delta SARS-CoV-2 genome reveals a single nucleotide mutation in the stem-loop II motif relative to ancestral SARS-CoV-2. Using unbiased molecular dynamics simulations, we found substantial differences in tertiary structure and dynamics relative to the earlier SARS-CoV-2 model. The characteristic 90 degree L-shaped kink did not form in the Delta structure, which instead adopted a more linear hairpin with limited bending dynamics. Increased fluctuation of the palindromic sequence in the terminal loop also produced an entropic penalty that helps explain reduced spontaneous homodimerization. Together, the structural and thermodynamic changes establish a mechanistic basis for experimentally observed differences in migration and dimer formation.
DOI and authorship note
https://doi.org/10.1021/acsphyschemau.3c00008
# Contributed equally and shared first authorship. * Corresponding author.
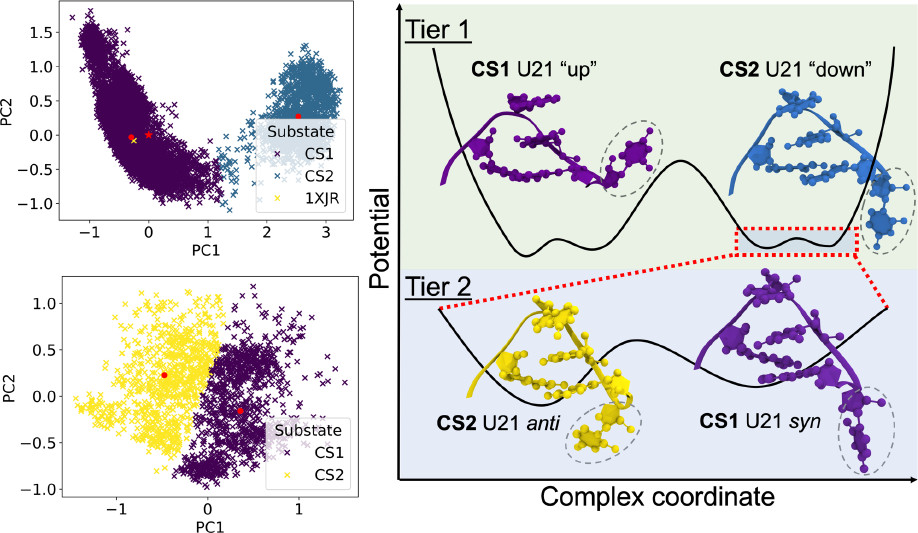
Structural, Dynamical, and Entropic Differences between SARS-CoV and SARS-CoV-2 s2m Elements Using Molecular Dynamics Simulations
Adam H. Kensinger#, Joseph A. Makowski#, Kendy A. Pellegrene#, Joshua A. Imperatore, Caylee L. Cunningham, Caleb J. Frye, Patrick E. Lackey, Mihaela Rita Mihailescu, Jeffrey D. Evanseck*
Abstract
The functional role of the highly conserved stem-loop II motif in SARS-CoV and SARS-CoV-2 remains enigmatic, and atomistic coordinates for the SARS-CoV-2 element were not available when this study began. We constructed three-dimensional coordinates consistent with NMR solution data and compared the resulting conformational ensembles with the SARS-CoV element. Molecular dynamics simulations showed that the SARS-CoV terminal pentaloop remains rigidly organized for possible kissing-dimer formation, whereas the SARS-CoV-2 terminal loop expands into a highly dynamic nonaloop. The resulting flexibility and structural disorganization introduce an entropic penalty that helps explain experimentally observed reductions in kissing-complex formation.
DOI and authorship note
https://doi.org/10.1021/acsphyschemau.2c00032
# Contributed equally and shared first authorship. * Corresponding author.
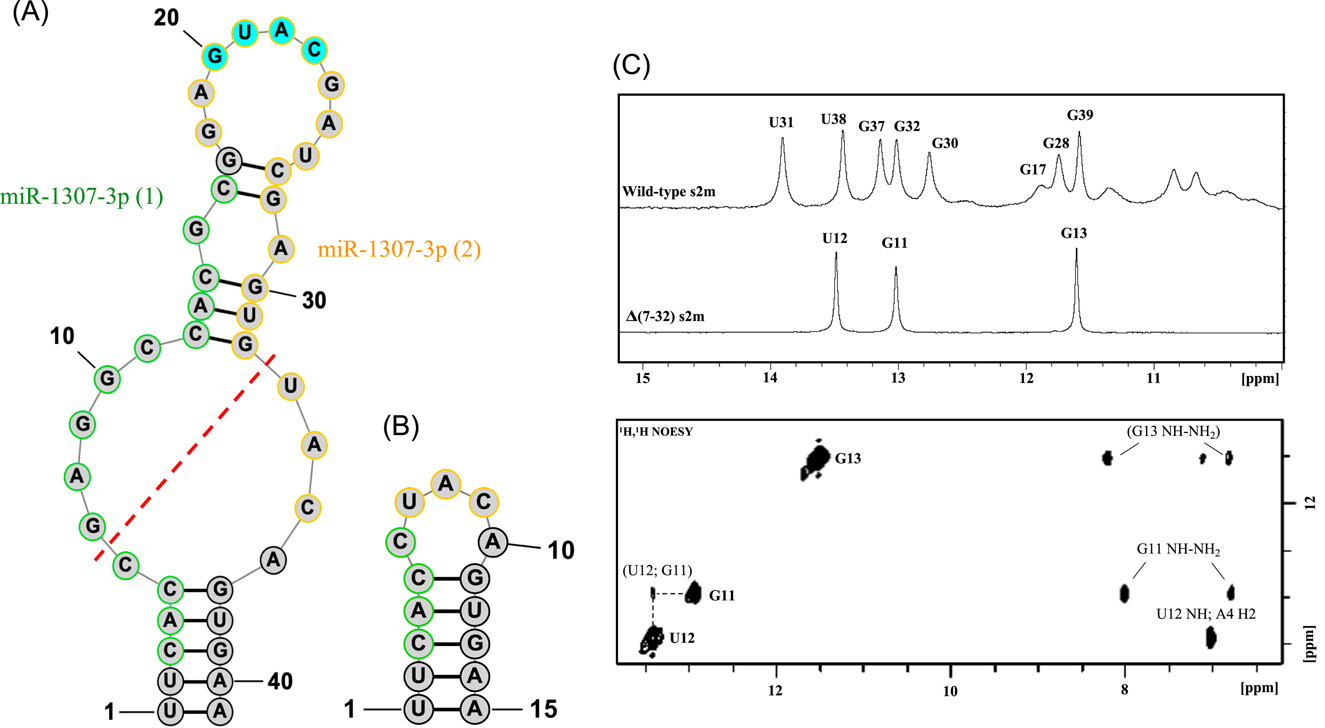
Bioinformatics analysis of the s2m mutations within the SARS-CoV-2 Omicron lineages
Caleb J. Frye, Morgan Shine, Joseph A. Makowski, Adam H. Kensinger, Caylee L. Cunningham, Ella J. Milback, Jeffrey D. Evanseck, Patrick E. Lackey, Mihaela Rita Mihailescu*
Abstract note
No formal abstract was provided on the journal page. Briefly, the paper reports a truncation of the s2m element in Omicron SARS-CoV-2 by bioinformatic analysis, offers experimental context, and discusses the possibility that the change may abolish microRNA binding in a way that benefits the virus.
DOI
https://doi.org/10.1002/jmv.28141
* Corresponding author noted on the publication.
Cover Art Gallery
Journal covers by my colleague Adam Kensinger.

ACS Physical Chemistry Au, Volume 3 Issue 5 cover art.

ACS Physical Chemistry Au, Volume 3 Issue 1 cover art.

Journal of Medical Virology cover art.